Part I: Panel suggestions for improving the CRISPR workflow
Following the insights gained from our The Market for CRISPR/Cas9 Genome Editing Products report (released June 2015 on the Interactive Market Intelligence platform), we fielded a follow-up questionnaire to our expert panel to quantify how the emergence of CRISPR is impacting the use of traditional tools and techniques.
The following insights are Part One of a series of releases that will provide even more insight into the evolving market of CRISPR technology. Our IMI clients will have access to the FULL report within their dashboard once the series has concluded.
This week, we are revealing some verbatim answers from the questionnaire.
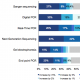
In the CRISPR IMI, our panel revealed that “validation of genome editing results” was the one area they would most like to improve. We asked 300+ global scientists currently using CRISPR technology how they would propose solving this, specifically:
“A majority of scientists said ‘validation of genome editing results’ was the one area they would most like to improve. How would you propose solving this problem?”
“Ensure the targeting is unique and specific. Southern blots to confirm that only specific targeting events have occurred using probes is required to ensure there are no other insertions. Otherwise, DNA-Seq to check whole genome is a reliable albeit expensive validation method.”
Staff Scientist
The National University of Singapore
“Good quality antibody reagents and good functional assays. Also, using other techniques such as site-directed mutagenesis and RNAi knockdown to understand the biology more fully.”
Senior Scientist
AstraZeneca UK
“The best way we have found to validate genome editing is to move to an in vivo system by SCNT and cloning. Further analysis and characterization of the genotype and phenotype of the resulting animals provide us with the most complete data sets.”
Senior Scientist
Recombinetics
“Sequencing to detect the point of insertion, similar to DNA modifications, enrich for guide RNA insertions with a probe and sequence fragments that are bound to the probe.”
Postdoctoral Fellow
Harvard Medical School
“I don’t see any obvious possible improvement. I think the design of the target should be the main focus to avoid off target effects which are much more difficult to identify.”
Postdoctoral Fellow
University of Queensland (Australia)
Want to see more verbatim insights from The Science Advisory Board? Let us know!
Next week: In Part Two of our “CRISPR Workflow” series, we will share quantitative data that answers how our panel believes feel that current tools, techniques and methods (eg. NGS, Sanger sequencing, transfection protocols) will be impacting due to the advent of CRISPR technology in their lab. Stay tuned!