The Next Generation of Next Generation Sequencing Technology
Last week, we highlighted key advantages of Portable Molecular Spectroscopy. While portable spectrometers make it possible to analyze unknown materials, analyzing unknown and unexplored information from DNA is just as crucial. Recently, scientists have been able to gather much information from our exons, the parts of DNA that are considered coding regions for proteins. However, there is still much to be understood about our genomes from the introns, or what until now has been considered “junk.” Maybe surprisingly, these “junk” regions may be of equal importance.
Today, I will delve into Next Generation Sequencing (NGS) technology for DNA — or rather, the next generation of NGS:
Perhaps the most interesting aspect of the laboratory instrumentation market is the evolution of technology. Next-generation sequencing, also referred to as high-throughput sequencing or second-generation sequencing,
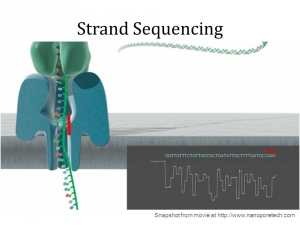
Nanopore Technology: Strand Sequencing from Oxford Nanopore’s 3rd Generation Sequencing Technology
has been a very exciting technology to observe — not only for how it has advanced discovery and influenced the life sciences market, but also because it is moving into the next phase of its technological maturation. In a sense, we are entering the next generation of Next-Generation sequencing.
New sequencing technologies are beginning to enter the market, ushering in the third-generation of DNA sequencing, also referred to as long-read sequencing. Second-generation sequencing methods require DNA to be fragmented into short lengths prior to sequencing, followed by re-assembly of sequences to piece together a genome. In contrast, long-read sequencing, capable of generating read lengths of 10kb or more, significantly reduces the challenge of whole genome assembly. This improves the ability to sequence complex genomes with repetitive regions, reducing ambiguity. Advances are also making sequencing more efficient, increasingly sensitive, and significantly lower in cost.
The first of the long-read technologies that have emerged commercially were nanopore technology (introduced
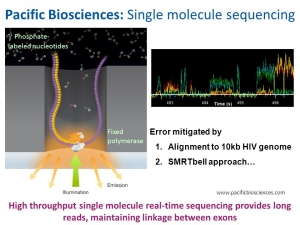
PacificBio’s single molecule real time sequencing
to the market by Oxford Nanopore), and single molecule real time sequencing (introduced to the market by Pacific Biosciences). Other methods are in various stages of development, with some expected to become commercially available within the forecast period of SDi’s 2018 Global Assessment report.
With the introduction of these new technologies, demand in the sequencing market is once again shifting. Over the past few years, demand for large-scale second-generation sequencers, which are capable of whole-genome sequencing and gene expression profiling, has declined as demand has stabilized and researchers turn to personal NGS systems for their targeted sequencing and analysis needs. However, this trend is already beginning to reverse, with positive demand growth for large-scale sequencers projected for 2019 and beyond.
Strong demand for large-scale sequencers will be driven by all sectors, but of note is the hospital and clinical industry. In 2017, Illumina promised that the cost of sequencing a human genome would soon be as low as $100. Though that price point has not yet been reached, and likely won’t be for several years, the plummeting cost of sequencing coupled with the rise of personalized medicine is sure to accelerate demand in the applied sector.
New competitors are poised to enter the market, potentially doubling the number of participants in the sequencers space. Several, including SeqLL, QuantuMDx, Stratos Genomics, and NabSys, are likely very close to bringing their systems to market. Of course, the development phase is perilous, and several companies that had previously been on SDi’s radar have seemingly ceased development or have gone out of business altogether.
These are very exciting times for molecular genetics. Keep up with the latest data in this ever-changing market with our 2018 Global Assessment report, the gold standard of market research for the life science and analytical instrument industries. Our next technology spotlight will cover how Triple Quadrupole ICP-MS is tackling food safety issues.